# if not alraedy done load library ggplot2
library(ggplot2)
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, uniondata(iris)
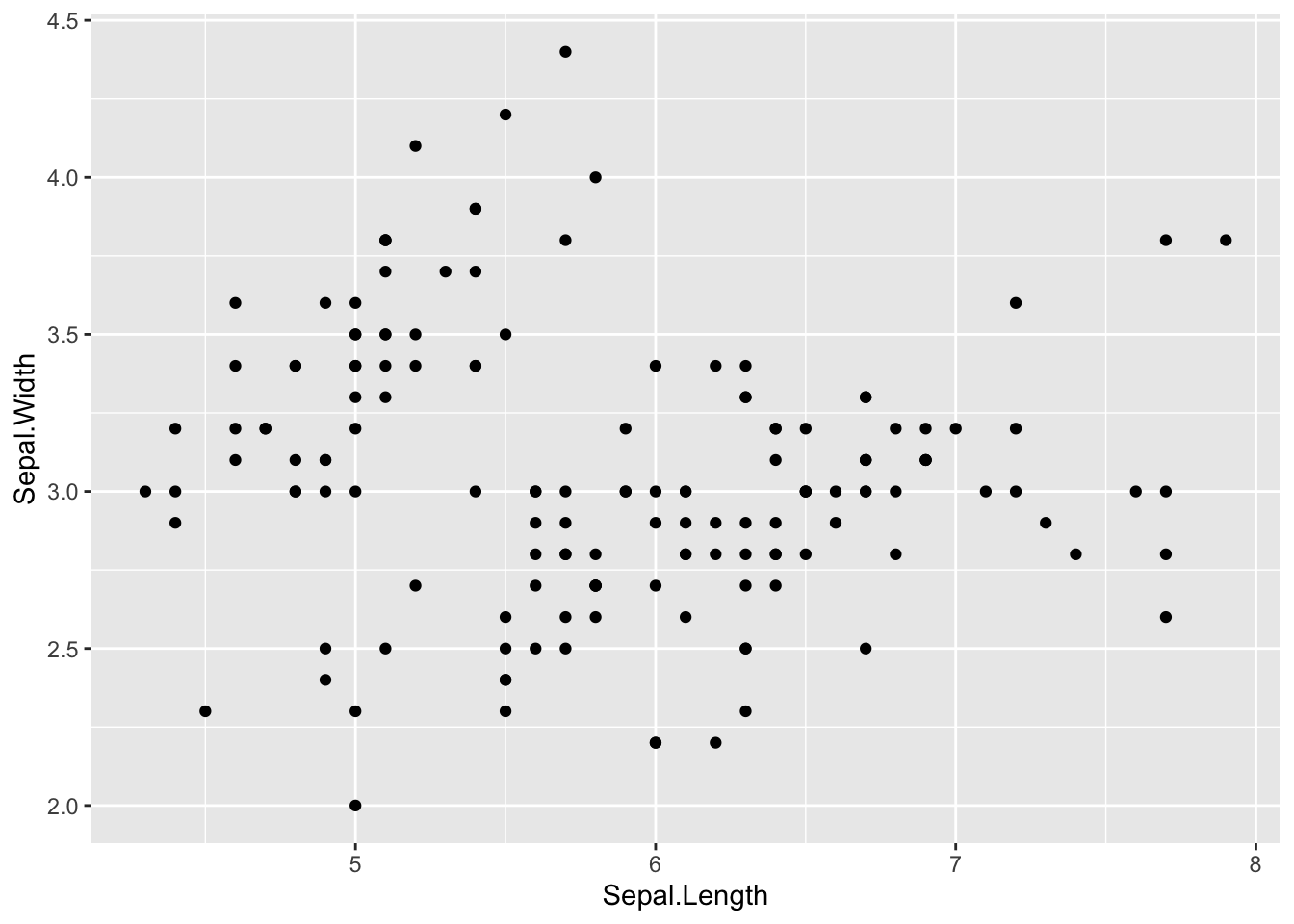
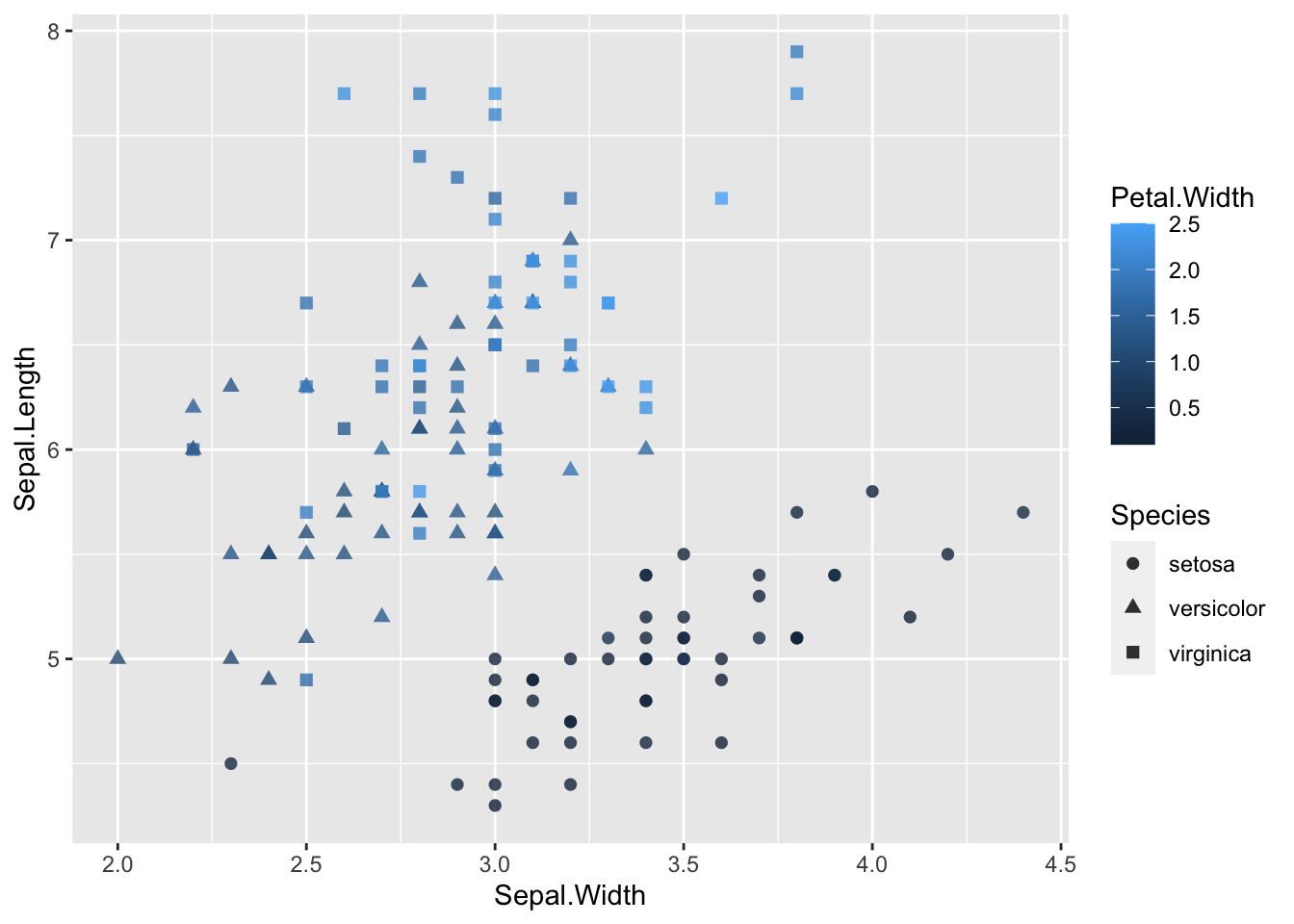
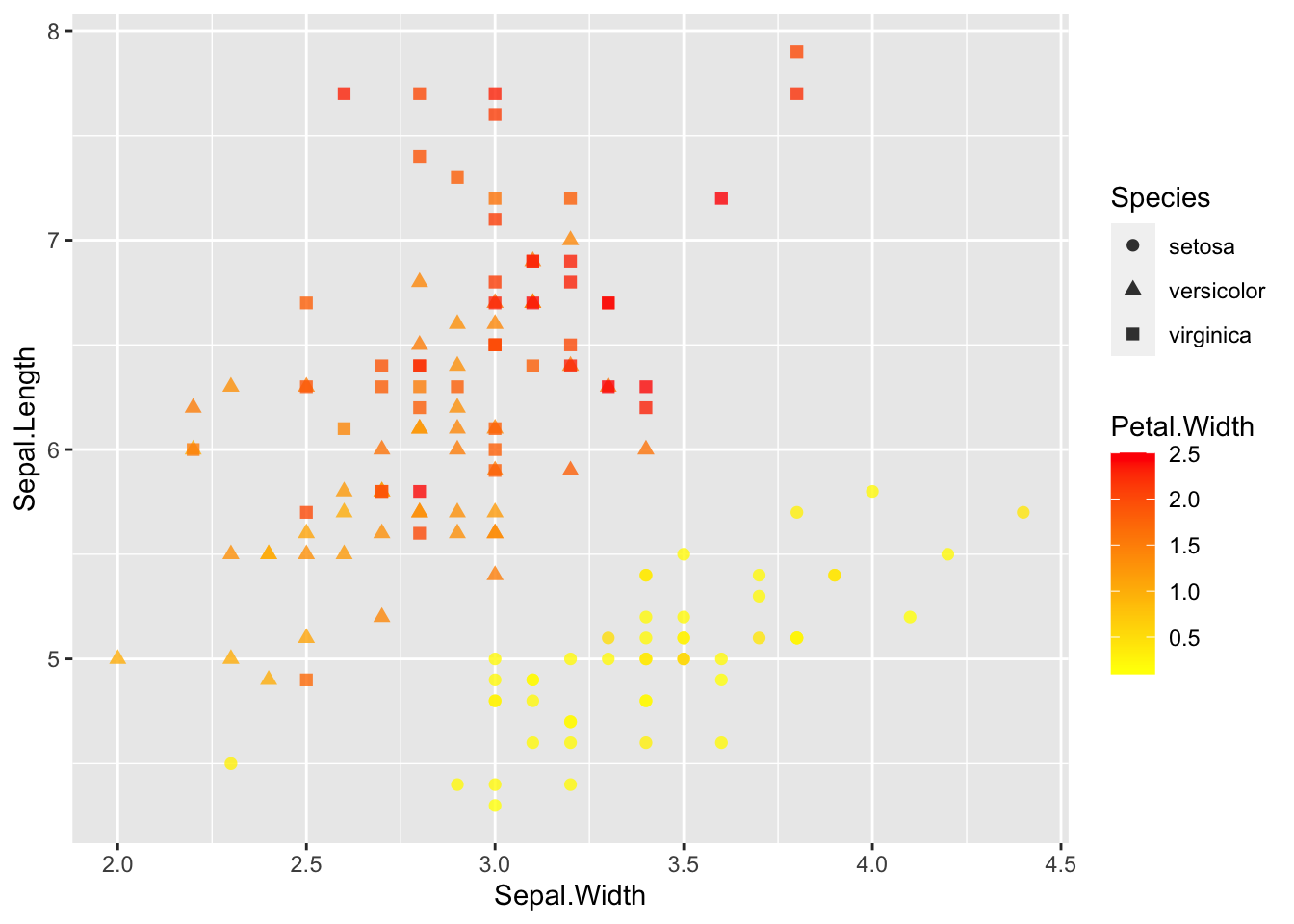
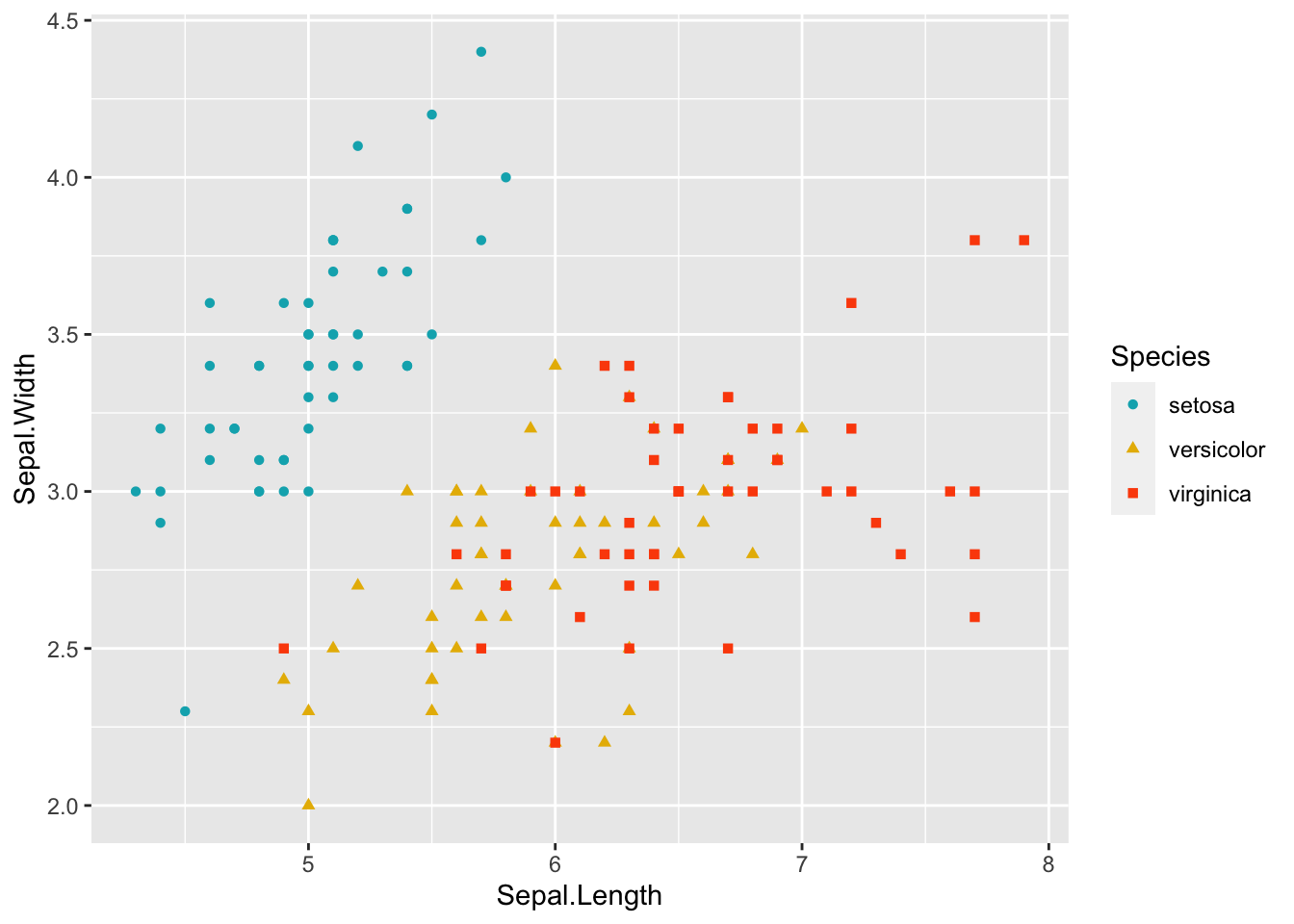
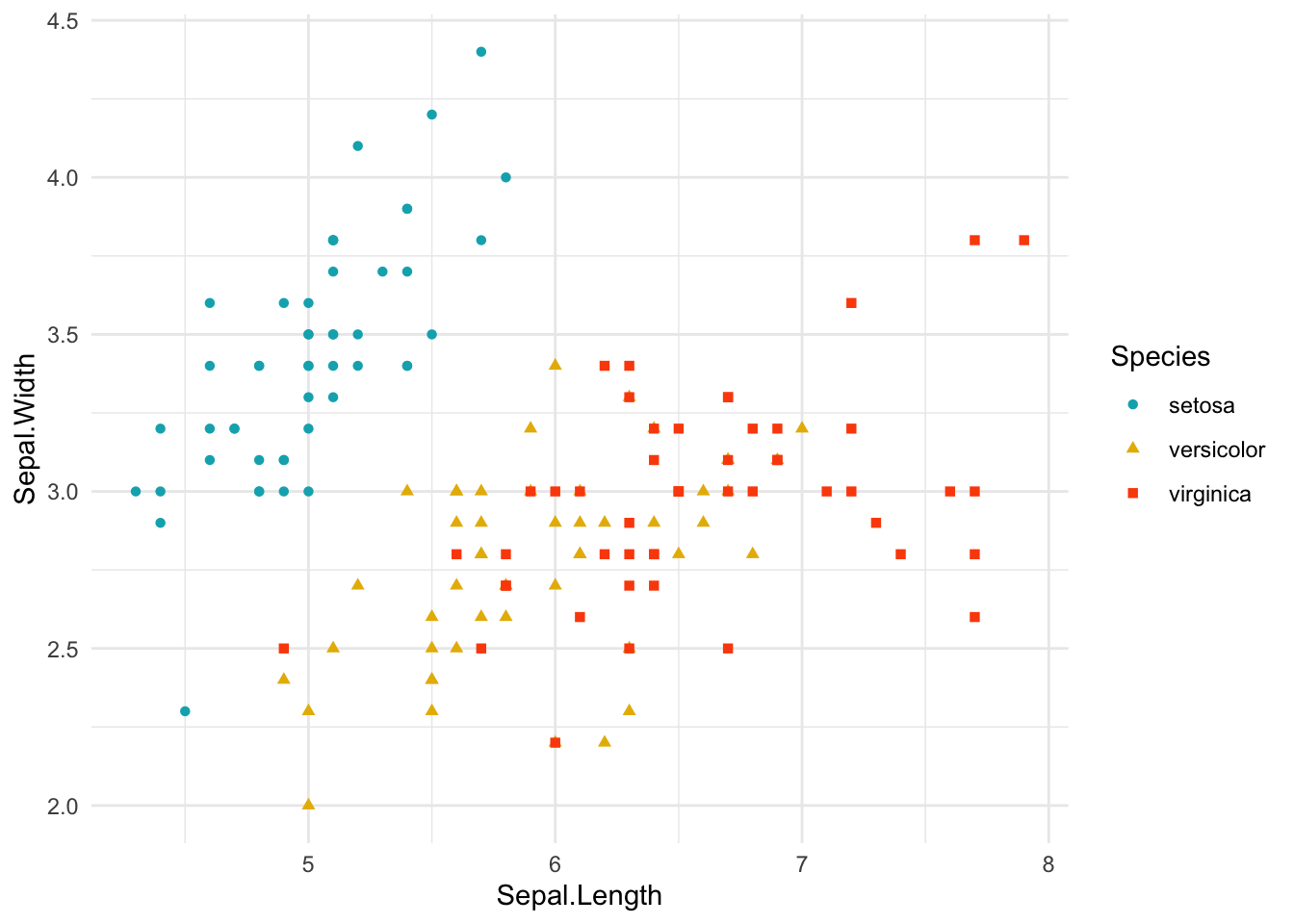
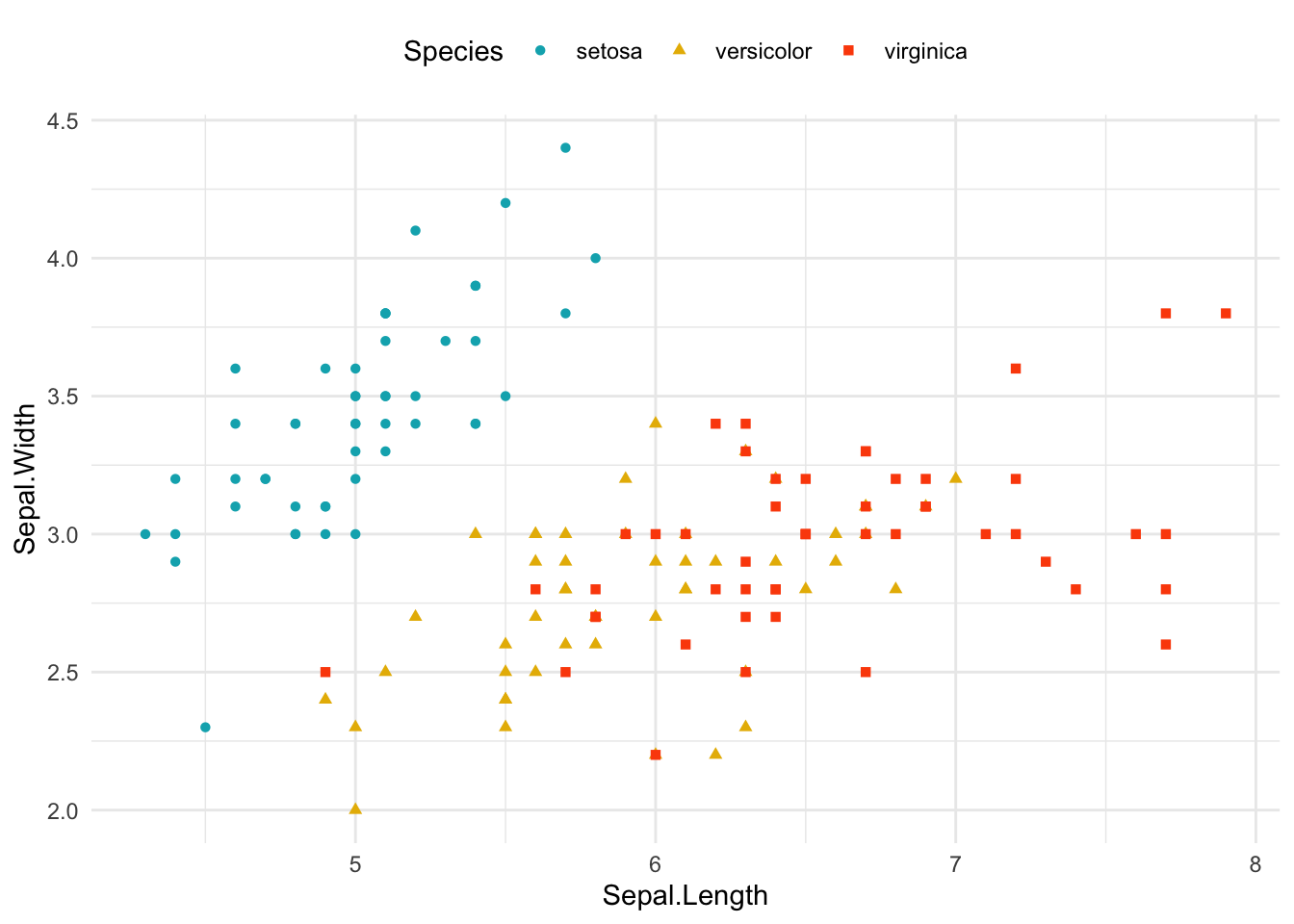
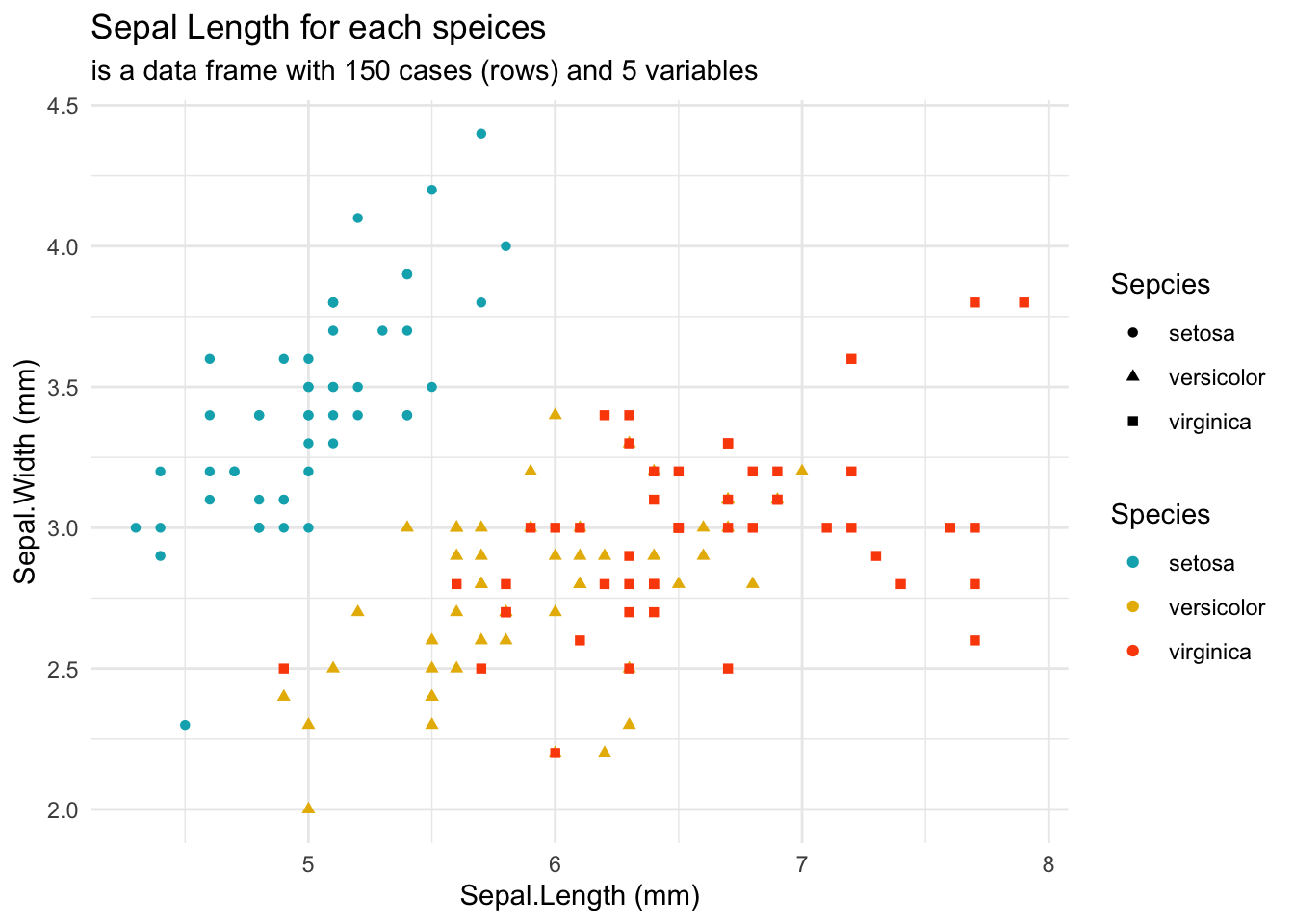
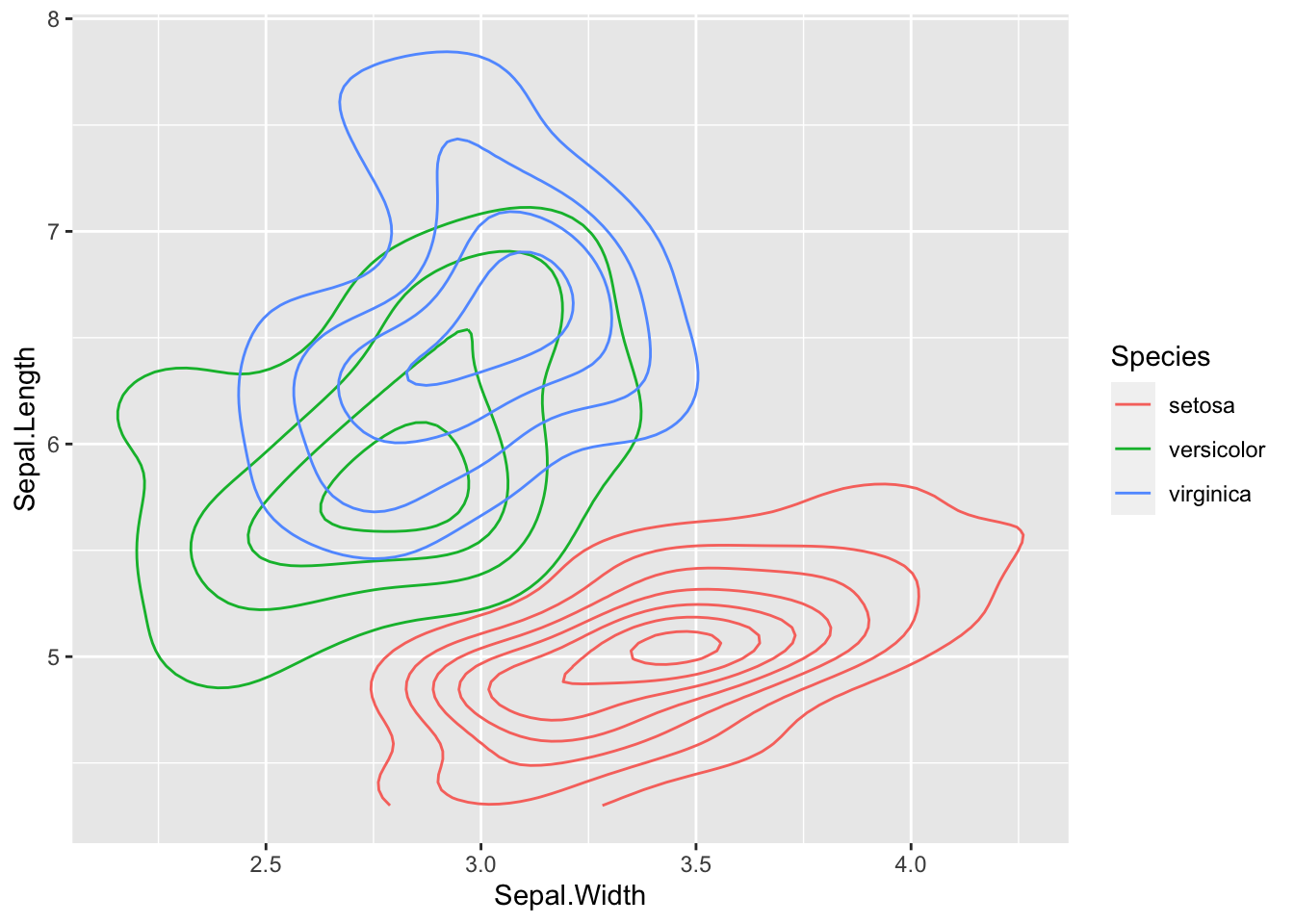
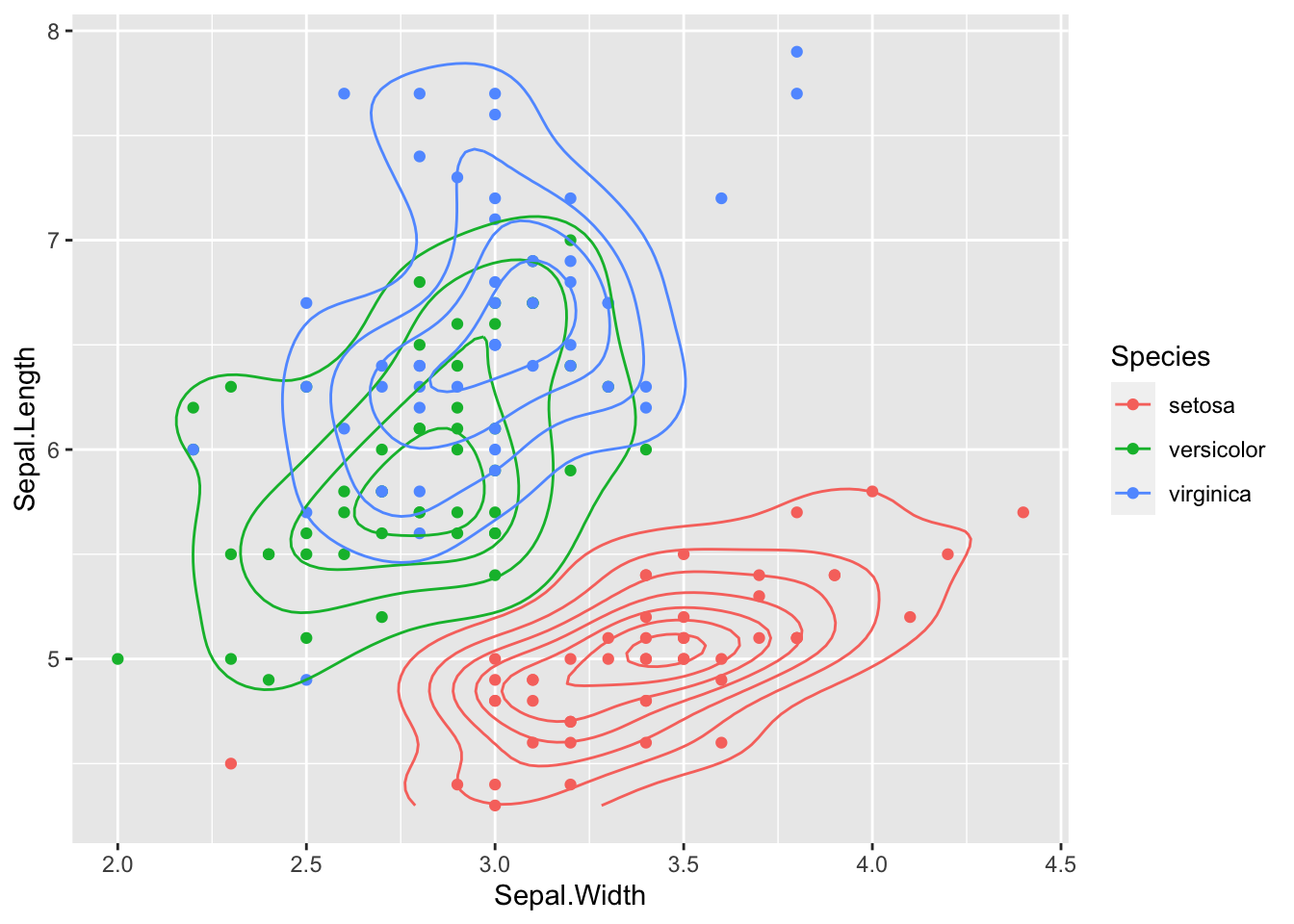
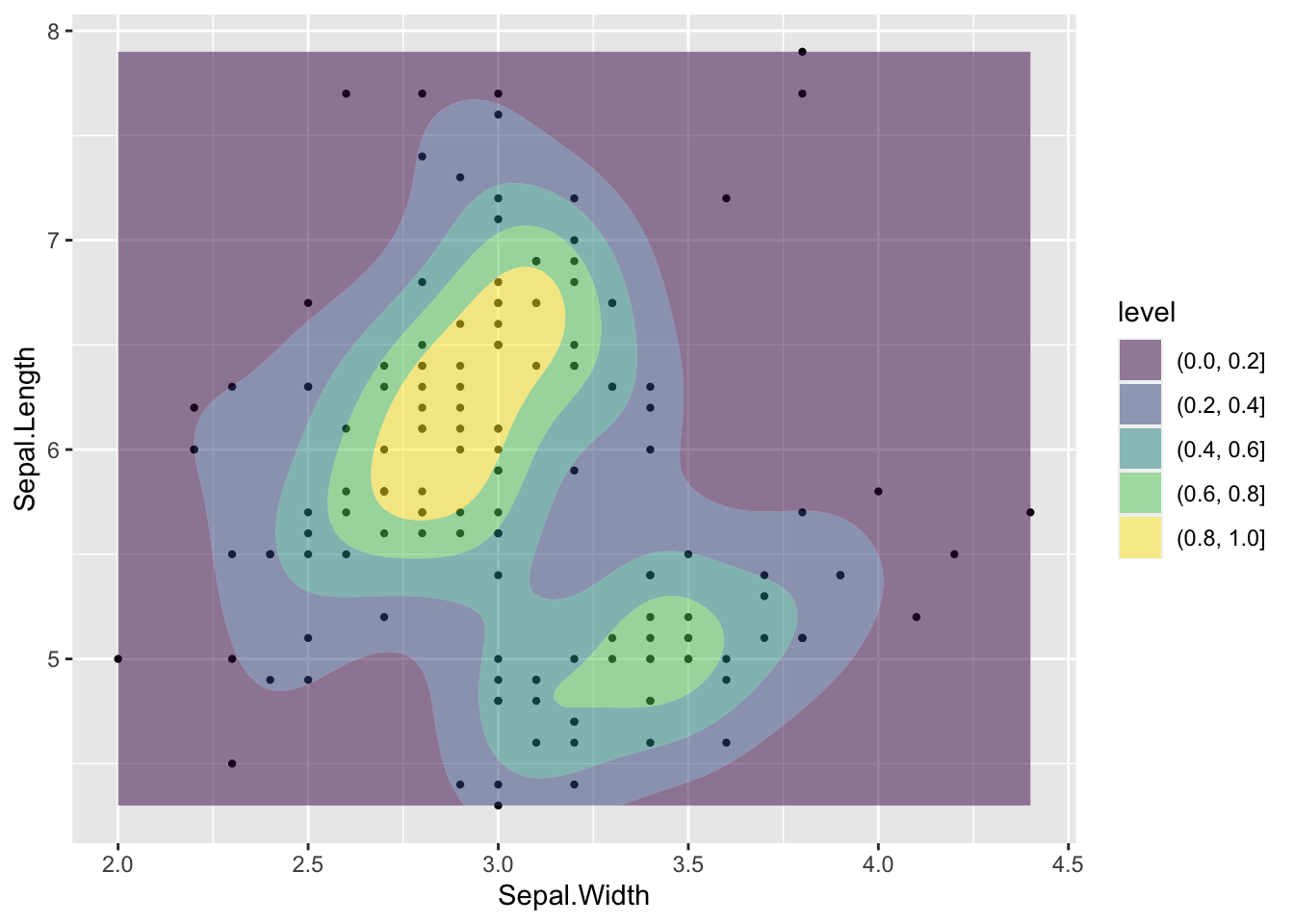
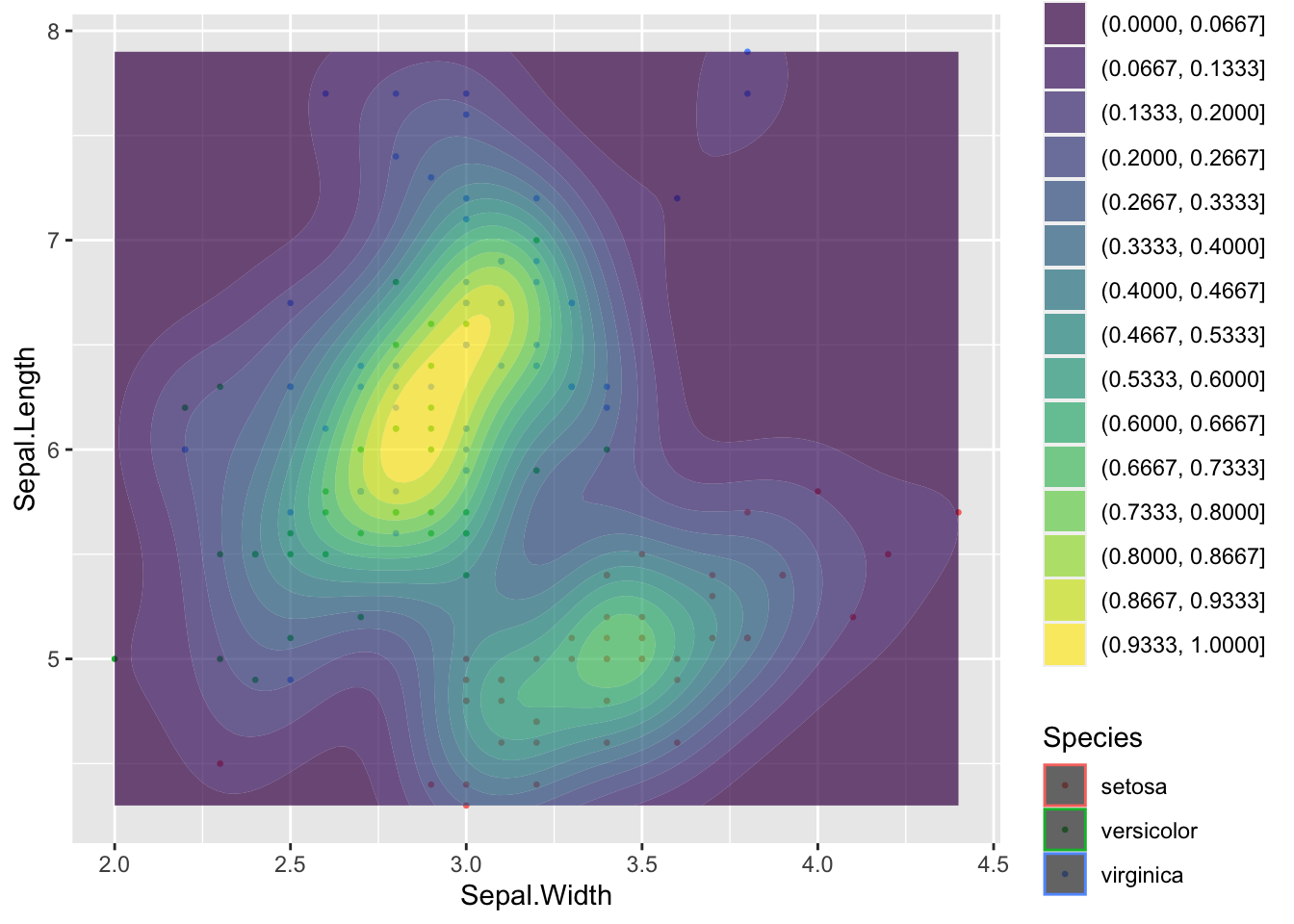
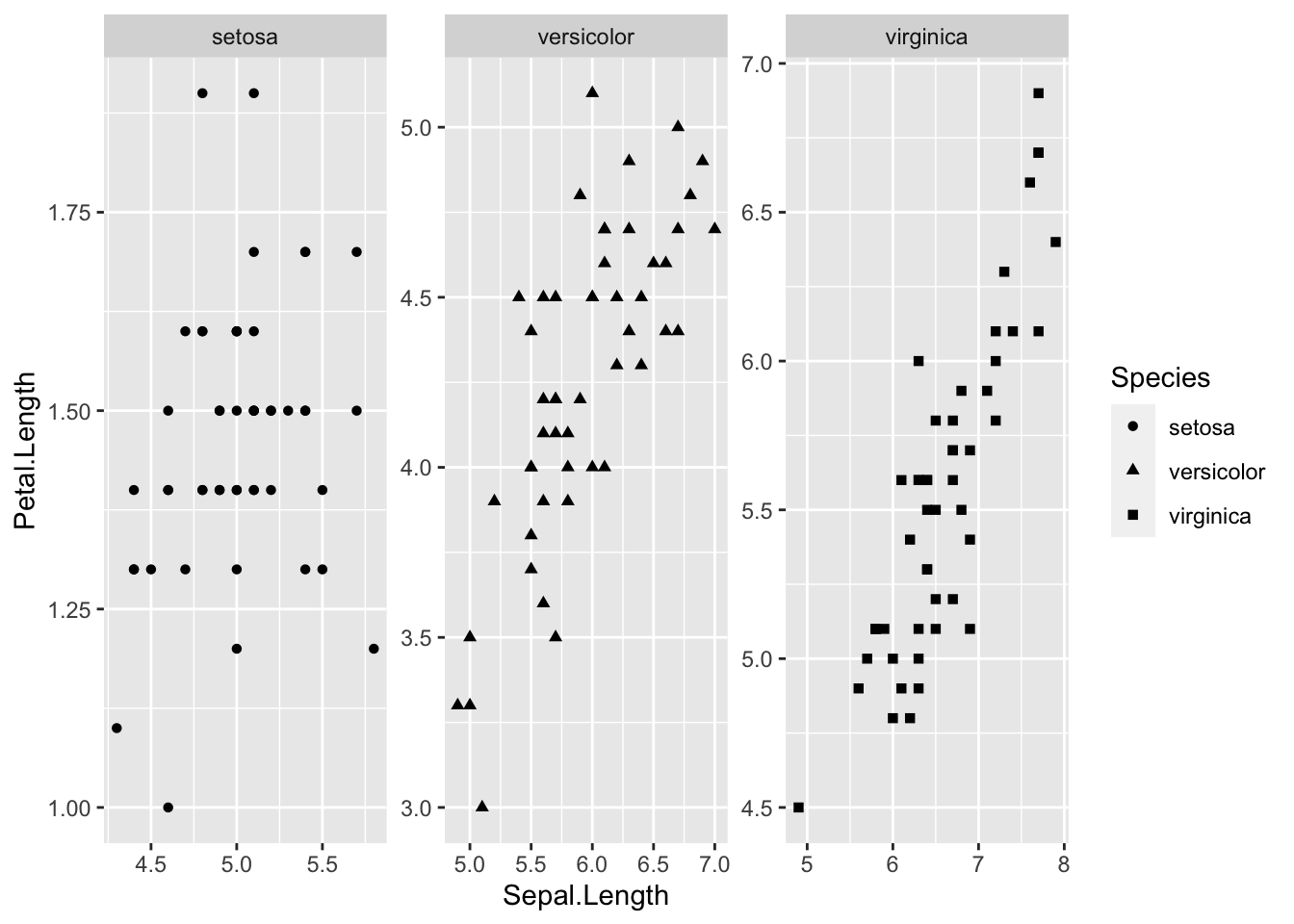
ggplot(data = iris, aes(Sepal.Length, Sepal.Width))+ # scatter plot
geom_point()